Nucleotide sequences from the EMBL Nucleotide Sequence Database, and protein sequences from UniProt (the Universal Protein Resource)ĮndNote is a popular reference and bibliography manager. Sequence files generated by the Mac program DNA Strider, containing one Nucleotide or Protein sequence. pro files are used in Lasergene, a sequence analysis tool produced by DNAStar. Files containing only metadata can be imported onto existing sequences in Geneious, see section 3.2.1ĭNAStar. For more information on importing primers from a spreadsheet, see the PCR Primers section. For files containing sequences, including nucleotides, proteins, primers or probes, Geneious will create a new document containing the sequence and any additional fields chosen for import. Sequences, primers and metadata information stored in spreadsheets can be uploaded to Geneious from either. csfasta files represent the color calls generated by the SOLiD sequencing system. HQ625581 MRVMGIPRNWPQWWIWGILGFWIMLMCRVEENSWVTVYYGVPVWKEATTTLFCASDAKAYĪBI. HQ625568 MRVRGTQRNWPQWWIWTSLGFWIILMCR-GNLWVTVYYGVPVWTDAKTTLFCASDAKAY 
HQ625588 MRVMGKWRNCQQWWIWGILGFWIILICN-AEQLWVTVYYGVPVWKEAKTTLFCASDAKAY HQ625572 MRVKGILKNYQQWWIWVILGFWMLMICNVVGNQWVTVYYGVPVWREAKATLFCASDAKAY HQ625589 MRVKGRSRNYPQWWVWGILGFWMFMICNGVGNRWVTVYYGVPVWKEAKATLFCASDAKAY HQ625570 MRVMGMWRNYPQWWIWGILGLWM-ICSVVGKLWVTVYYGVPVWTDAKATLFCASDAKAY An example Clustal file:ĬLUSTAL W (1.74) multiple sequence alignment Ĭlustal format files are used to store multiple sequence alignments and contain the word clustal at the beginning. The Clustal format is used by the well known multiple sequence alignment programs ClustalW, ClustalX and Clustal Omega. pd4 are not currently supported for import. Currently it does not import other fields, restriction cut sites or primer binding sites. This will import name, description, topology, sequence and annotations. Geneious can import annotated sequences files in the standard Clone Manager molecule format. 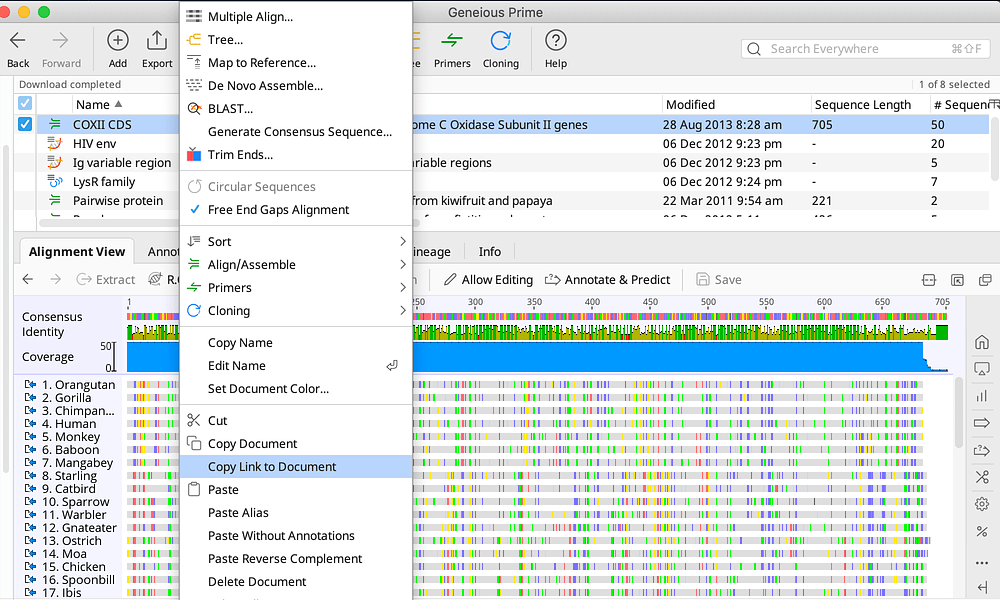
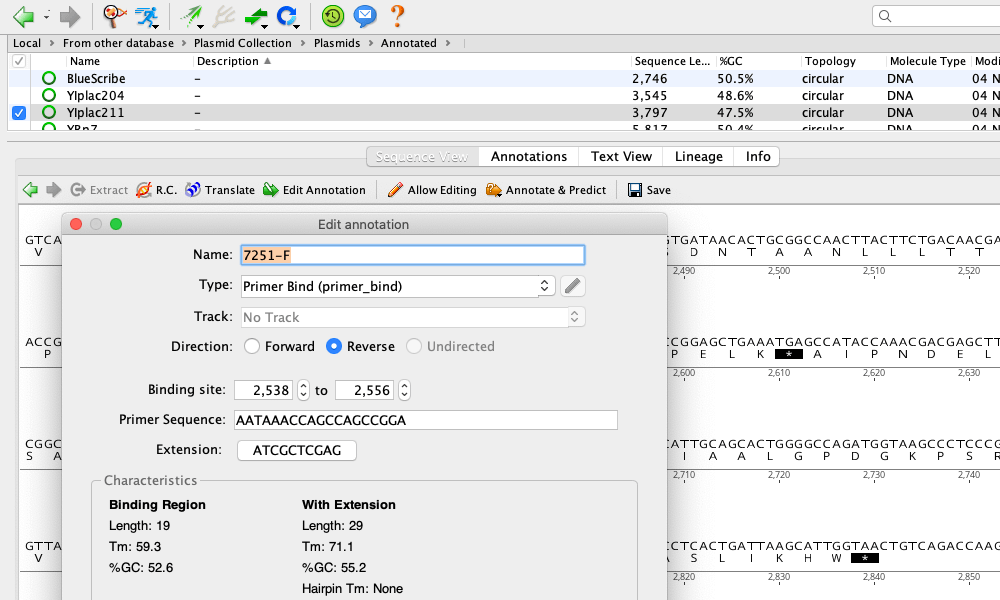
You can use a BED file to annotate existing sequences in your local database, import entirely new sequences, or import the annotations onto blank sequences. The BED format contains sequence annotation information. Matching Geneious document fields to your spreadsheetģ.2.2 Importing Vector NTI Databases BED annotations To export a folder from the Local, simply select the folder and in the Toolbar click on “Export”, “Export Folder…”, choose the file destination and name, and click “Save”.CSV/TSV (Comma/Tab Separated Values) and Excel spreadsheet filesģ.2.1 Importing metadata from a spreadsheet onto existing documents Fill in as much information as possible for future reference.Ī very convenient tool is the ability to export and import folders of primers. Here, various metadata such as Gene, Organism, Direction etc. Highlight the primer in the Document Table, and go to the “Info” tab in the Document Viewer. Once a primer has been created, any associated information can be edited. Here, enter the primer sequence, name, and in the “Type” dropdown menu select “Primer”. To add a new primer, highlight the destination folder, then in the Menu Bar select “Sequence” followed by “New Sequence” from the dropdown menu. From the Toolbar, click “Add” then “New folder…” and enter the new folder name. To create a new folder in the directory, highlight the local folder in the Sources Panel. If a primer folder does not exist in the Local Directory, one should be made. SI Barcode Network Informatics Documentation
0 Comments
Leave a Reply. |

 RSS Feed
RSS Feed
